Interpretable Deep Learning for Regulatory Genomics
The Koo Lab studies how regulatory information is encoded in biological sequences and how this information can be learned, interpreted, and stress tested using modern machine learning. Deep learning models now achieve strong predictive performance across a wide range of regulatory genomics tasks, yet their scientific value depends on whether they can support mechanistic insight rather than prediction alone. Our work is guided by the view that interpretability is not a post hoc visualization problem, but a property that must be evaluated, audited, and often designed for.
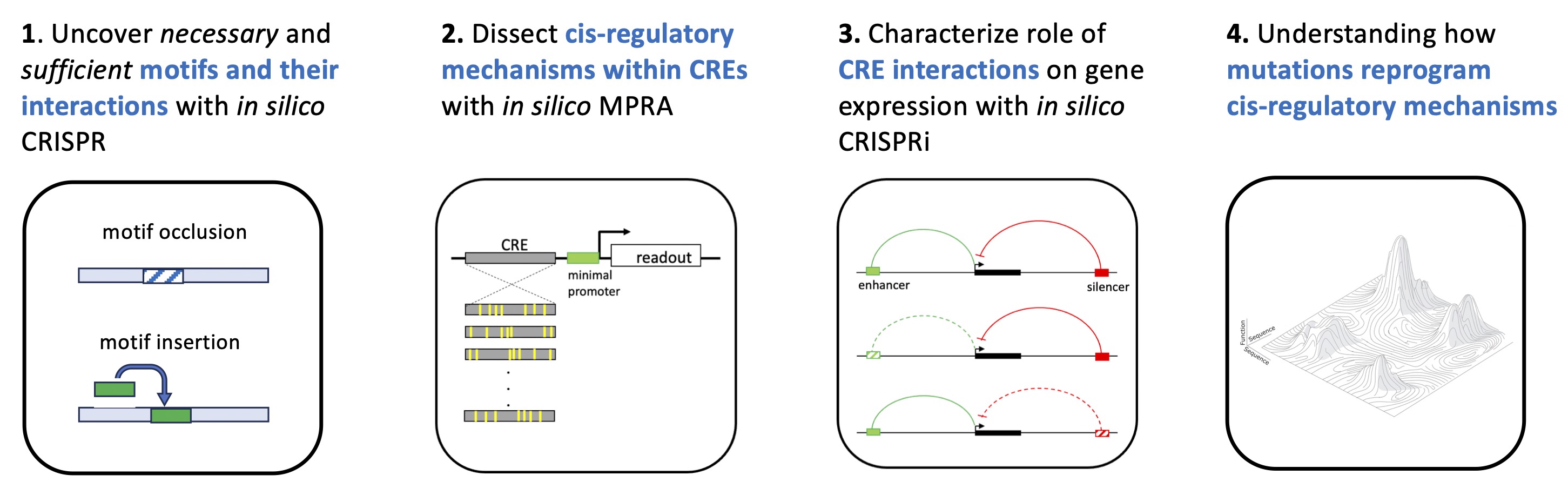
We develop methods to probe what sequence features and contextual dependencies deep models rely on, with a particular emphasis on regulatory DNA. This includes frameworks for attribution analysis, counterfactual perturbations, and synthetic sequence experiments that expose model behavior beyond natural genomic variation. A recurring theme in our work is that standard attribution methods can be misleading if applied uncritically, especially in the presence of model uncertainty, distribution shift, or correlated sequence features. As a result, we focus on developing principled tests for explanation reliability and on connecting model-derived features to experimentally testable hypotheses.
More broadly, we are interested in understanding how architectural choices, training objectives, and data regimes influence what models learn about regulatory logic. Rather than treating deep networks as black boxes, we aim to characterize their inductive biases, identify failure modes, and establish when their predictions and explanations can be trusted. This philosophy extends across our work in regulatory genomics, cancer genomics, and related sequence modeling problems.
Generative Models for Regulatory Sequence Design
A central focus of the lab is the development of generative models that can design biological sequences conditioned on regulatory function. Unlike traditional predictive models, which estimate function for a fixed sequence, generative models explicitly learn distributions over sequences and offer a direct way to explore and manipulate regulatory space.
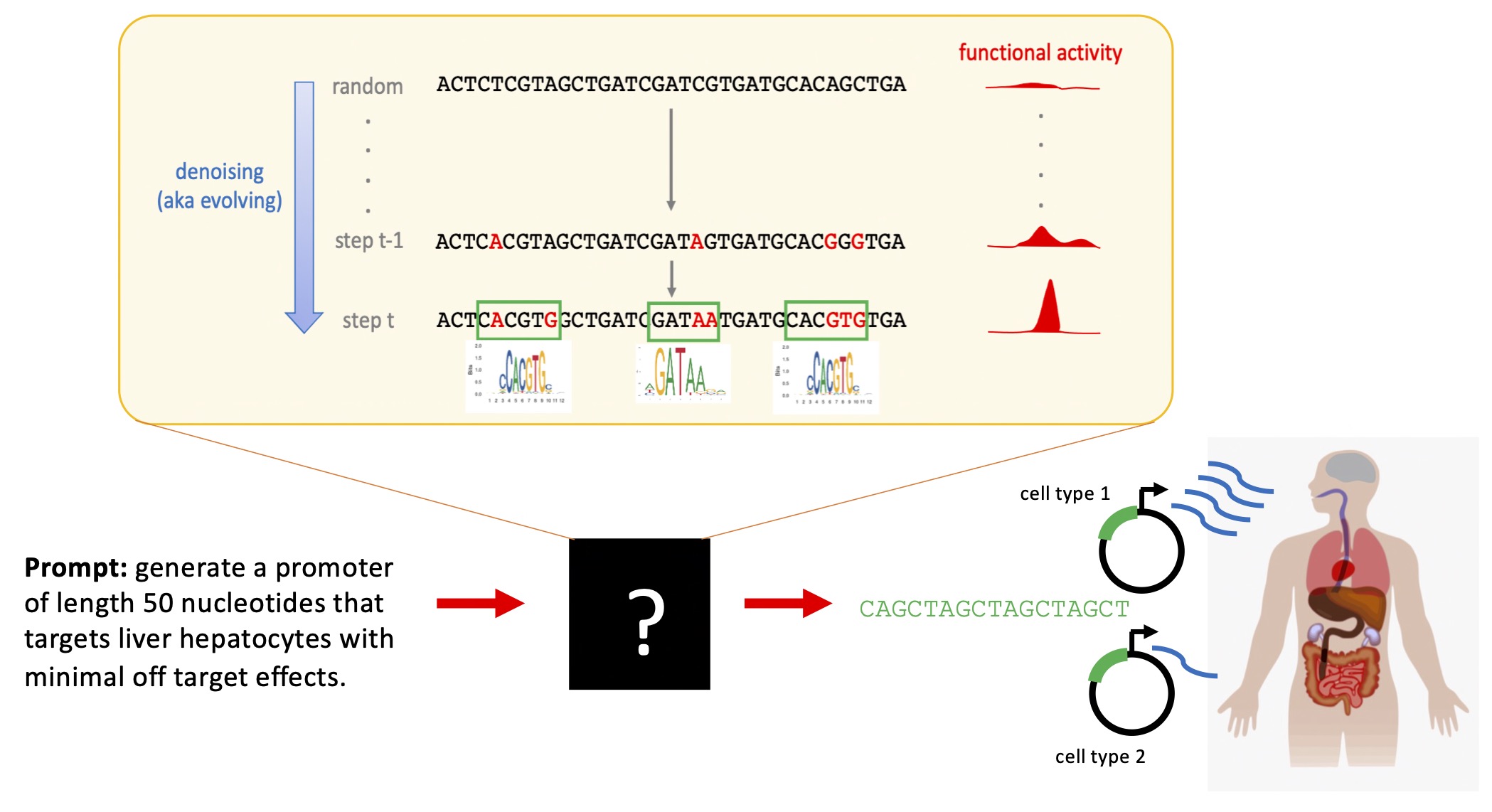
We have developed discrete diffusion-based generative models for DNA that operate directly on symbolic sequences and are conditioned on cell type specific or context specific regulatory signals. These models are designed to address challenges that arise in regulatory genomics, including sparse motif structure, strong class imbalance, and the need to generate diverse yet functional sequences. By integrating supervision directly into the generative process, these models provide a framework for designing sequences that satisfy complex regulatory constraints rather than optimizing a single score.
Beyond sequence generation, we use these models as scientific tools to study what aspects of regulatory logic are captured by different training objectives and data sources. This includes evaluating whether generated sequences reflect known regulatory principles, whether they generalize across genetic backgrounds, and how they compare to alternative generative approaches. A key goal is to move toward generative models that are not only useful for design, but also interpretable and experimentally falsifiable.
Improving Generalization through Active and Continual Learning
A persistent limitation of current genomic models is their fragility under distribution shift. Models trained on specific cell types, genetic backgrounds, or experimental assays often fail when applied outside their training domain, even when predictions appear confident. Our work addresses this limitation by studying generalization as a first class problem rather than a secondary benchmark.
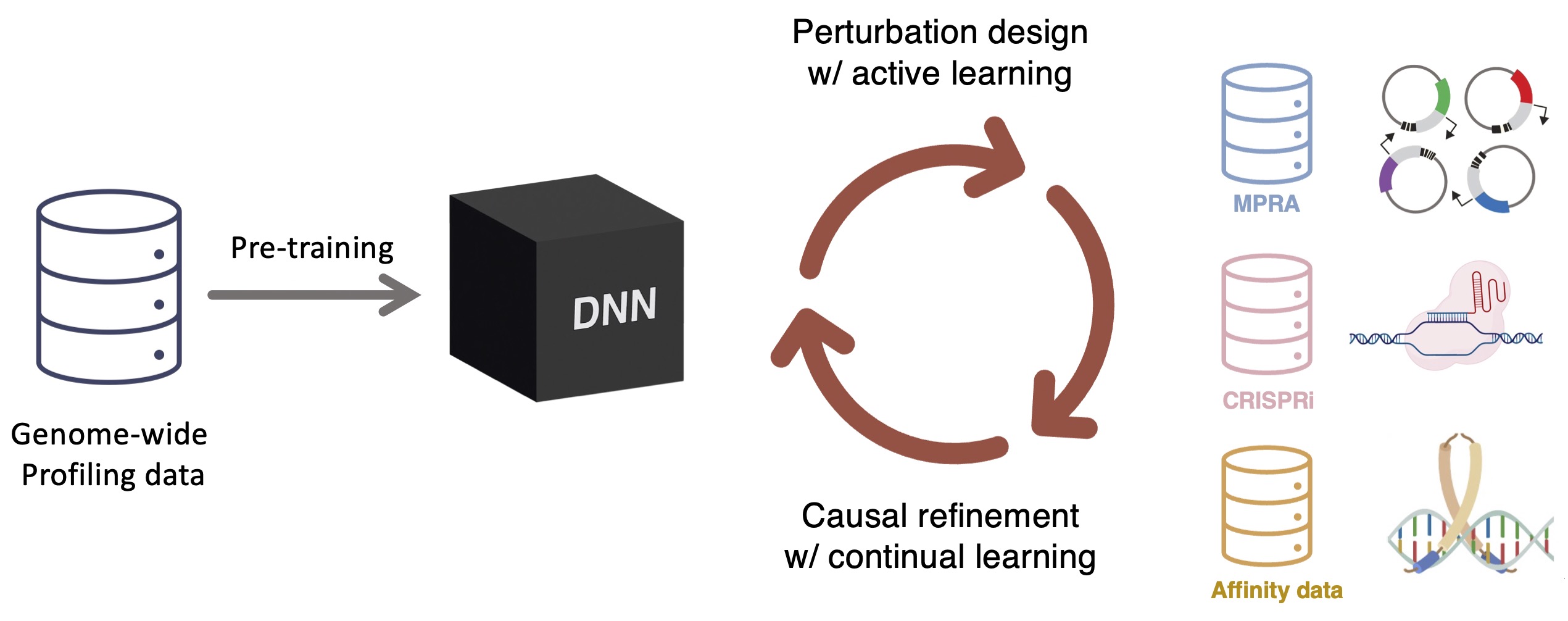
We are developing active and continual learning frameworks that integrate modeling with targeted experimental perturbations. In this setting, models are not trained once on a static dataset, but are iteratively refined using new data chosen to be maximally informative. This includes selecting sequences or perturbations that expose uncertainty, challenge model assumptions, or probe underrepresented regions of sequence space. By closing the loop between prediction, uncertainty estimation, and data acquisition, we aim to build models that improve systematically rather than overfit passively collected data.
We are also interested in how knowledge learned in one biological context can be transferred to new systems with limited data. This includes adapting models across cell types, species, and experimental modalities, as well as understanding when such transfer fails. Ultimately, our goal is to develop genomic AI systems that are robust, self correcting, and capable of supporting discovery in settings where data are scarce and biological complexity is high.
Integration with Experimental Genomics
A central component of the Koo Lab's research is its integration with the Quantitative Biology experimental core facility at Cold Spring Harbor Laboratory, led by Ken Chang. The core generates functional genomics datasets that are closely aligned with model development, interpretation, and validation. This tight coordination enables low latency cycles between computational analysis and experimental execution, allowing modeling results to rapidly inform experimental design and newly generated data to be incorporated into training and evaluation. The close coordination between modeling and experimentation enables rapid iteration and supports investigation of regulatory mechanisms.